What is PCA
PCA(Principal Component Analysis) is a non-parametric linear method to reduce dimension. It’s a very popular way used in dimension reduction.
How to use it
Use PCA is much easier than understand it. So, I put the function first. If we look at the document of sklearning, it says like this:
pca = PCA(n_components=2) # select how many components we want to pick up
pca.fit(X) # fit the PCA model, or we can use fit_transform(X)
...
pca.transform(X2) # transfer data by trained model
We can also use line graph to plot the variance% that PCA explained.
variance = covar_matrix.explained_variance_ratio_ #calculate variance ratios
var=np.cumsum(np.round(covar_matrix.explained_variance_ratio_, decimals=3)*100)
plt.ylabel('% Variance Explained')
plt.xlabel('# of Features')
plt.title('PCA Analysis')
plt.ylim(30,100.5)
plt.style.context('seaborn-whitegrid')
plt.plot(var)

Don’t forget nomalization before PCA
How does it work?
1. First, let us see what our data looks like

We call left distribution as low redundancy, and right distribution as high redundancy. In the case of high redundancy, there are many related dimensions, e.g., how many hours you start and how many score you get in the test. PCA is used to reduce redundancy and noise in the dataset, and find the strongest signal.
2. How could we find signal and Nosie?
The answer is covariance matrix.
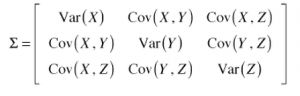
Diagonal is variance of x,y,z(3 dimensions) which is signal, off diagonal is covariance which is redundancy.
3. How could we increase signal and decrease redundancy?
Simple! make covariance matrix as a diagonal matrix, like this:

We need transfer original dataset X to dataset Y through linear transformation P: PX = Y, which makes covariance matrix of Y: Sy to be a diagonal matrix.We can seem this to be a changing the basis of coordinate, in the new coordinate, we can get the maxium variance in one axis.
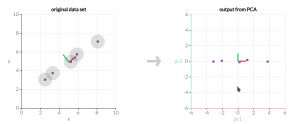
More inforamtion and 3-D simulation, please visit: http://setosa.io/ev/principal-component-analysis/
4. Math approvel
Sy = 1⁄(n-1) YYT
Sy = 1⁄(n-1)(PX)(PX)T
Sy = 1⁄(n-1)(PXXTPT)
Sy = 1⁄(n-1)(P(XXT)PT)
Let A = XXT which is a symmetric matrix, Sy = 1⁄(n-1)(PAPT)
According to the characters of symmetric matrix, A can be written as VDVT, where D is diagonal matrix, V is eigenvector matrix.
Here is the tricky part: let P = V, then
Sy = 1⁄(n-1)(PAPT) = 1⁄(n-1)(PPTDPPT). since the inverse of orthonormal matrix is its transpose, P−1 = PT.
Sy = 1⁄(n-1)(PP-1DPP-1) = 1⁄(n-1) D. Here is what we want! D is a diagonal matrix!
So, V(eigenvectors) is our result for P.
5. How gets the eigenvectors?
You can find more information here. Basically, the blew is the formula: ![]()
Reference: A Tutorial on Data Reduction, Shireen Elhabian and Aly Farag
Leave a Reply