Spark provides spark MLlib for machine learning in a scalable environment. MLlib includes three major parts: Transformer, Estimator and Pipeline. Essentially, transformer takes a dataframe as an input and returns a new data frame with more columns. Most featurization tasks are transformer. Estimator takes a dataframes as an input and returns a model(transformer), as we know the ML algorithms.. Pipeline combines transformer and estimator together.
Additionally, data frame becomes the primary API for MLlib. There is not any more new features for RDD based API in Spark MLib.
If you already understood or used high level machine learning or deep learning frameworks, like scikit-learning, keras, tersorflow, you will find everything is so familiar with. But when you use spark MLlib in practice, you still need third library’s help. I will talk about it in the end.
Basic Stats
# Corrlation
from pyspark.ml.stat import Correlation
r1 = Correlation.corr(df, "features").head()
print("Pearson correlation matrix:\n" + str(r1[0]))
# Summarizer
from pyspark.ml.stat import Summarizer
# compute statistics for multiple metrics without weight
df.select(summarizer.summary(df.features)).show(truncate=False)
# ChiSquare
r = ChiSquareTest.test(df, "features", "label").head()
print("pValues: " + str(r.pValues))
print("degreesOfFreedom: " + str(r.degreesOfFreedom))
print("statistics: " + str(r.statistics))Featurization
# TF-IDF
# stop word remove
remover = StopWordsRemover(inputCol="raw", outputCol="filtered")
remover.transform(sentenceData)
# tokenize
tokenizer = Tokenizer(inputCol="sentence", outputCol="words")
wordsData = tokenizer.transform(sentenceData)
# n grame
ngram = NGram(n=2, inputCol="wordsData", outputCol="ngrams")
ngramDataFrame = ngram.transform(wordDataFrame)
# word frequence
hashingTF = HashingTF(inputCol="words", outputCol="rawFeatures", numFeatures=20)
featurizedData = hashingTF.transform(wordsData)
# idf
idf = IDF(inputCol="rawFeatures", outputCol="features")
idfModel = idf.fit(featurizedData)
rescaledData = idfModel.transform(featurizedData)
# word2vec
word2Vec = Word2Vec(vectorSize=N, minCount=0, inputCol="text", outputCol="result")
model = word2Vec.fit(documentDF)
# binarizer
binarizer = Binarizer(threshold=0.5, inputCol="feature", outputCol="binarized_feature")
binarizedDataFrame = binarizer.transform(continuousDataFrame)
# PCA
# reduce dimension to 3
pca = PCA(k=3, inputCol="features", outputCol="pcaFeatures")
model = pca.fit(df)
# StringIndex
# encodes a string column of labels to a column of label of indices order by frequency or alphabet
indexer = StringIndexer(inputCol="category", outputCol="categoryIndex")
indexed = indexer.fit(df).transform(df)
# OneHotEstimator
# we need to use StringIndex first if apply to categorical feature
encoder = OneHotEncoderEstimator(inputCols=["categoryIndex1", "categoryIndex2"],
outputCols=["categoryVec1", "categoryVec2"])
model = encoder.fit(df)encoded = model.transform(df)
# Normalize & Scaler
normalizer = Normalizer(inputCol="features", outputCol="normFeatures", p=1.0)
lInfNormData = normalizer.transform(dataFrame, {normalizer.p: float("inf")})
l1NormData = normalizer.transform(dataFrame)
# standard scaler
# withMean=false: standard deviation, withMean=true: mean
scaler = StandardScaler(inputCol="features", outputCol="scaledFeatures",
withStd=True, withMean=False)
# maxmin scaler
scaler = MinMaxScaler(inputCol="features", outputCol="scaledFeatures")
# max abs scaler
scaler = MaxAbsScaler(inputCol="features", outputCol="scaledFeatures")
# bin
from pyspark.ml.feature import Bucketizer
splits = [-float("inf"), -0.5, 0.0, 0.5, float("inf")]
bucketizer = Bucketizer(splits=splits, inputCol="features", outputCol="bucketedFeatures")
# QuantileDiscretizer
discretizer = QuantileDiscretizer(numBuckets=3, inputCol="hour", outputCol="result")
# ElementwiseProduct
transformer = ElementwiseProduct(scalingVec=Vectors.dense([0.0, 1.0, 2.0]),
inputCol="vector", outputCol="transformedVector")
# SQL Transformer
sqlTrans = SQLTransformer(
statement="SELECT *, (v1 + v2) AS v3, (v1 * v2) AS v4 FROM __THIS__")
# VectorAssembler
# combine vector together for future model inputs
assembler = VectorAssembler(
inputCols=["hour", "mobile", "userFeatures"], # the columns we need to combine
outputCol="features") # output column
# Imputer
# handle missing value
imputer = Imputer(inputCols=["a", "b"], outputCols=["out_a", "out_b"])
imputer.setMissingValue(custom_value)
# slice vector
slicer = VectorSlicer(inputCol="userFeatures", outputCol="features", indices=[1])
# ChiSqSelector
# use Chisqare to select the features
selector = ChiSqSelector(numTopFeatures=1, featuresCol="features",
outputCol="selectedFeatures", labelCol="clicked")Clarification and Regression
# Linear regression
lr = LinearRegression(maxIter=10, regParam=0.3, elasticNetParam=0.8)
# logistic regression
from pyspark.ml.classification import LogisticRegression
lr = LogisticRegression(maxIter=10, regParam=0.3, elasticNetParam=0.8)
mlr = LogisticRegression(maxIter=10, regParam=0.3, elasticNetParam=0.8, family="multinomial") # multinomial
# decision tree
# classification
labelIndexer = StringIndexer(inputCol="label", outputCol="indexedLabel").fit(data)
featureIndexer =\
VectorIndexer(inputCol="features", outputCol="indexedFeatures", maxCategories=4).fit(data)
(trainingData, testData) = data.randomSplit([0.7, 0.3])
dt = DecisionTreeClassifier(labelCol="indexedLabel", featuresCol="indexedFeatures")
# regression
dt = DecisionTreeRegressor(featuresCol="indexedFeatures")
# Random forest
# classification
rf = RandomForestClassifier(labelCol="indexedLabel", featuresCol="indexedFeatures", numTrees=10)
# regeression
rf = RandomForestRegressor(featuresCol="indexedFeatures")
# gradient-boosted
gbt = GBTRegressor(featuresCol="indexedFeatures", maxIter=10)
# preceptron
trainer = MultilayerPerceptronClassifier(maxIter=100, layers=layers, blockSize=128, seed=1234)
# SVM
# Linear SVM, there is no kernel SVM like RBF
lsvc = LinearSVC(maxIter=10, regParam=0.1)
# Naive Bayes
nb = NaiveBayes(smoothing=1.0, modelType="multinomial")
# KNN
knn = KNNClassifier().setTopTreeSize(training.count().toInt / 500).setK(10)Clustering
# k-means
kmeans = KMeans().setK(2).setSeed(1)
model = kmeans.fit(dataset)
# GMM
gmm = GaussianMixture().setK(2).setSeed(538009335)Collaborative Filtering
als = ALS(maxIter=5, regParam=0.01, userCol="userId", itemCol="movieId", ratingCol="rating",
coldStartStrategy="drop")
model = als.fit(training)
# Evaluate the model by computing the RMSE on the test datapredictions = model.transform(test)evaluator = RegressionEvaluator(metricName="rmse", labelCol="rating",predictionCol="prediction")
rmse = evaluator.evaluate(predictions)Validation
# split train and test
train, test = data.randomSplit([0.9, 0.1], seed=12345)
# cross validation
crossval = CrossValidator(estimator=pipeline,
estimatorParamMaps=paramGrid,
evaluator=BinaryClassificationEvaluator(),
numFolds=2) # use 3+ folds in practiceYou might be already found the problem. The ecosystem of Spark MLlib is not as rich as scikit learning, and it is lack of deep learning (of course, its name is machine learning). According to Databricks documents, We still have the solutions.
- Use scikit learning on single node. Very simple solution. but since scikit leanring load the data still in memory. If the note is faster enough(driver), we can get a good performance as well.
- To solve deep learning problem. we have two work around methods.
- Apply keras, tensorflow on single node with GPU acceleration(Recommend by databricks).
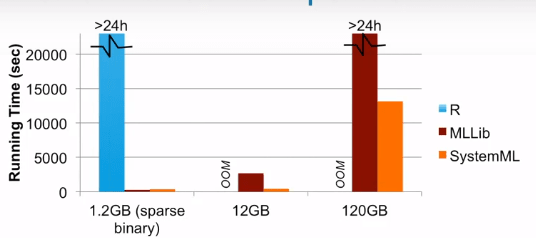
- Distribute Training. It might be slower than on the single node because of communication overhead. There are two frameworks used for distribute training. Horovod and Apache SystemML. I’ve never use Horovod, but you can find information here. As to SystemML, it is more like a wrapper for high level API and provide cluster optimizer which parses the code into spark RDD(live variable analysis, propagate stats, rewrite by matrix decomposition and runtime instruction). From the official website, we know it is much faster than MLLib and native R. The problem is it didn’t update anymore since 2017.
# Create and save a single-hidden-layer Keras model for binary classification# NOTE: In a typical workflow, we'd train the model before exporting it to disk,# but we skip that step here for brevity
model = Sequential()
model.add(Dense(units=20, input_shape=[num_features], activation='relu'))
model.add(Dense(units=1, activation='sigmoid'))
model_path = "/tmp/simple-binary-classification"
model.save(model_path)
transformer = KerasTransformer(inputCol="features", outputCol="predictions", modelFile=model_path)It seems no perfect solution for machine learning in spark, right? Don’t forget we have other time costing jobs: hyper parameters configuration and validation. We can run same model with different hyper parameters on different nodes using paramMap which similar to grid search or random search.
from pyspark.ml.tuning import CrossValidator, ParamGridBuilder
paramGrid = ParamGridBuilder().addGrid(lr.maxIter, [10, 100, 1000]).addGrid(lr.regParam, [0.1, 0.01]).build()
crossval = CrossValidator(estimator=pipeline,
estimatorParamMaps=paramGrid,
evaluator=RegressionEvaluator(),
numFolds=2) # use 3+ folds in practice
cvModel = crossval.fit(training)There might be someone saying: why we don’t use MPI? The answer is simple, too complex. Although it can gain the perfect performance, and you can do whatever you want even running distributed GPU + CPU codes, there are too many things we need to manually configuration on low level API without fail tolerance.
In conclusion, we can utilize spark for ELT and training/ validation model to maximize the performance(it did really well for these works). But until now, we still need third frameworks to help us do deep learning or machine learning tasks on single strong node.
Reference:
Spark MLlib: http://spark.apache.org/docs/latest/ml-guide.html
Databrick: https://docs.databricks.com/getting-started/index.html
Spark ML Tuning: http://spark.apache.org/docs/latest/ml-tuning.html
Harovod: https://github.com/horovod/horovod
Apache SystemML: https://systemml.apache.org/docs/1.1.0/beginners-guide-keras2dml
Leave a Reply